Machine Learning for Anatomical imaging
Machine learning can enhance MR and CT imaging through various means such as denoising, low-dose reconstruction, and task-based reconstruction.
Unsupervised learning for medical image denoising without high-quality images
Unsupervised learning for medical image denoising without high-quality images
Currently most neural network-based medical image denoising methods require matched or unmatched high-quality images as reference during training, which are inaccessible under certain circumstances such as dynamic imaging. Based on the recently proposed Noise2noise method, we proposed to train denoising network from noisy measurement only by splitting the measurement into two sets and mapping the reconstructed images from one set to the other. Theoretical analysis was done which indicated the feasible conditions of the Noise2noise scheme. We further proposed a novel consensus loss function based on the theory to better utilize both noisy measurements.
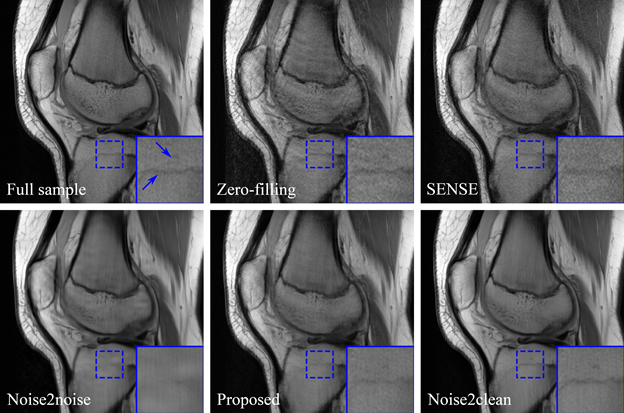
Wu, D., Gong, K., Kim, K. and Li, Q., 2019. Consensus Neural Network for Medical Imaging Denoising with Only Noisy Training Samples. arXiv preprint arXiv:1906.03639.
Iterative Low-dose CT reconstruction with unsupervised manifold learning via k-sparse autoencoder
Iterative Low-dose CT reconstruction with unsupervised manifold learning via k-sparse autoencoder
A k-sparse autoencoder was built to learn the manifold of normal-dose CT images in an unsupervised manner. Then objective function was built concerning both data fidelity and distance to the manifold. The objective function was solved via alternative optimization of the data fidelity part and the manifold part, which was a monotonic algorithm given appropriate step size. K-sparse autoencoder was built on patches and trained greedily by keeping only the k largest encodes at each training iteration. Furthermore, the k-sparse autoencoder demonstrated superior edge preservation performance compared to normal autoencoders and L1-sparse autoencoder.
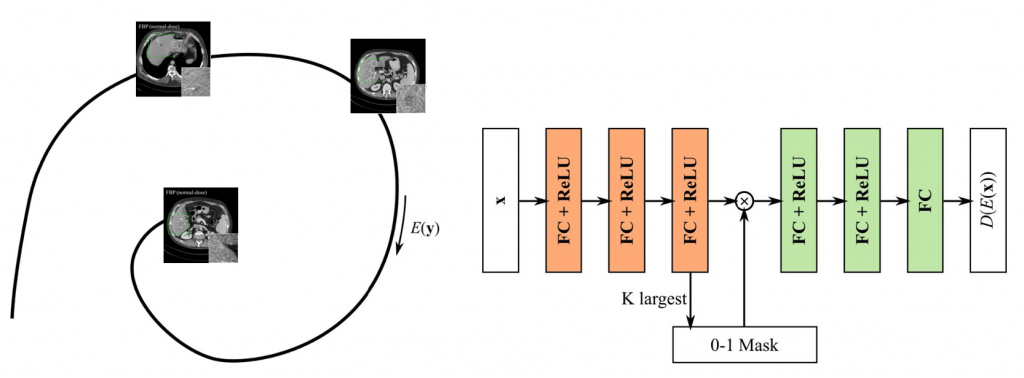
Wu, D., Kim, K. and Li, Q., 2019. Computationally Efficient Deep Neural Network for Computed Tomography Image Reconstruction. Medical physics.
Undersampled image reconstruction with computational efficient neural network
Undersampled image reconstruction with computational efficient neural network
One way to do image reconstruction with deep neural networks is unrolling iterative algorithms to finite steps and parameterize prior-related terms with trainable neural networks. By taking the measurement into consideration, the unroll approach achieved promising performance in undersampled problems such as few-view, limited-angle and low-dose. However, it requires significant amount of memory and training time for 3D volume reconstruction which is not practical for most current hardware. We proposed a computationally efficient training approach which broke the learning into sequential image-domain trainings and significantly reduced the computational cost. The estimated memory cost was reduced from 417 GB to 2 GB for a CT reconstruction problem with ordinal-size image (640*640*128). It was also demonstrated the proposed method could achieve similar performance to unrolled network.
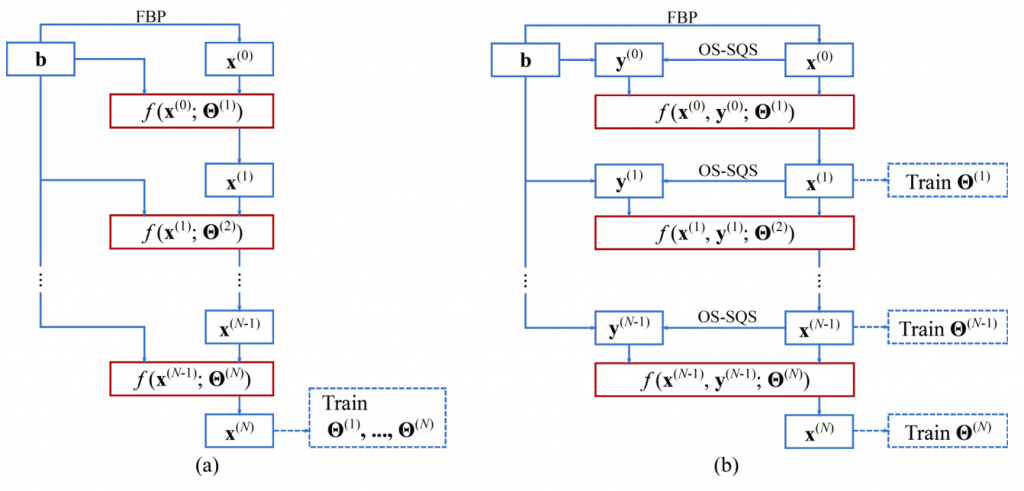
Wu, D., Kim, K. and Li, Q., 2019. Computationally Efficient Deep Neural Network for Computed Tomography Image Reconstruction. Medical physics.
Low-dose dual energy CT reconstruction via deep neural networks
Low-dose dual energy CT reconstruction via deep neural networks
Dual energy CT (DECT) is generally not feasible for very large patients due to the heavily attenuated photon flux under 80kVp. This project investigated the deep neural network-based image reconstruction for low-flux scenarios of DECT. Normal-dose and half-dose DECT scans were acquired for each of 40 patients under IRB approval. Noise was inserted to the normal-dose raw data to simulate low-dose scans and a learned primal-dual network was trained with L2 norm on 30 patients. The network was tested on the rest 10 patients’ real half-dose scans and achieved significantly reduced noise without bias.
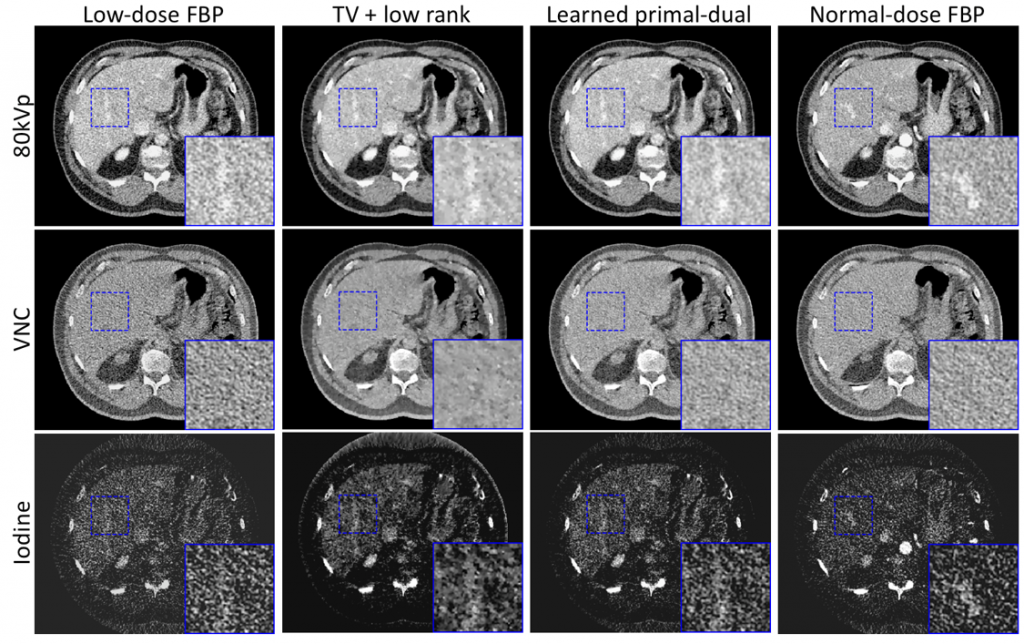
Wu, D., Kim, K., Kalra, M.K., De Man, B. and Li, Q., 2019, May. Learned primal-dual reconstruction for dual energy computed tomography with reduced dose. In 15th International Meeting on Fully Three-Dimensional Image Reconstruction in Radiology and Nuclear Medicine (Vol. 11072, p. 1107206). International Society for Optics and Photonics.
Task-based image reconstruction for lung nodule detection
Task-based image reconstruction for lung nodule detection
Current image reconstruction is optimized for visual quality of the images to radiologist. However, computer aided diagnosis system could rely on very different features for automatic lesion detection than human observers. We built a lung nodule detection network whose input was the sinogram rather than the images. The deep neural network consisted of a learned primal-dual reconstruction network and 3D CNN detection network. Improved FROC performance was achieved with the task-based image reconstruction on simulated sparse-view sinograms compared to reconstruction optimized for L2 norm loss on images.
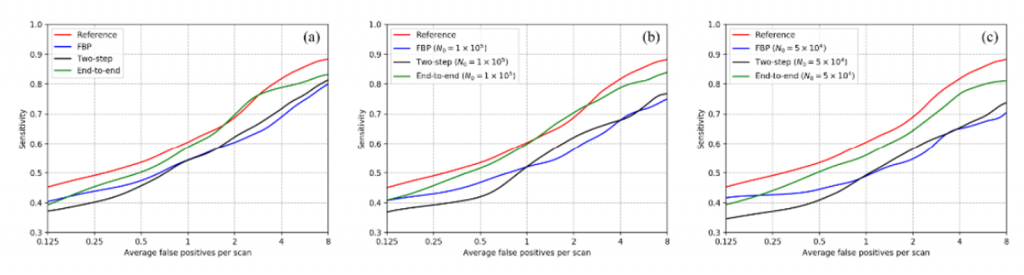
Wu, D., Kim, K., Dong, B., El Fakhri, G. and Li, Q., 2018, September. End-to-end lung nodule detection in computed tomography. In International Workshop on Machine Learning in Medical Imaging (pp. 37-45). Springer, Cham.

